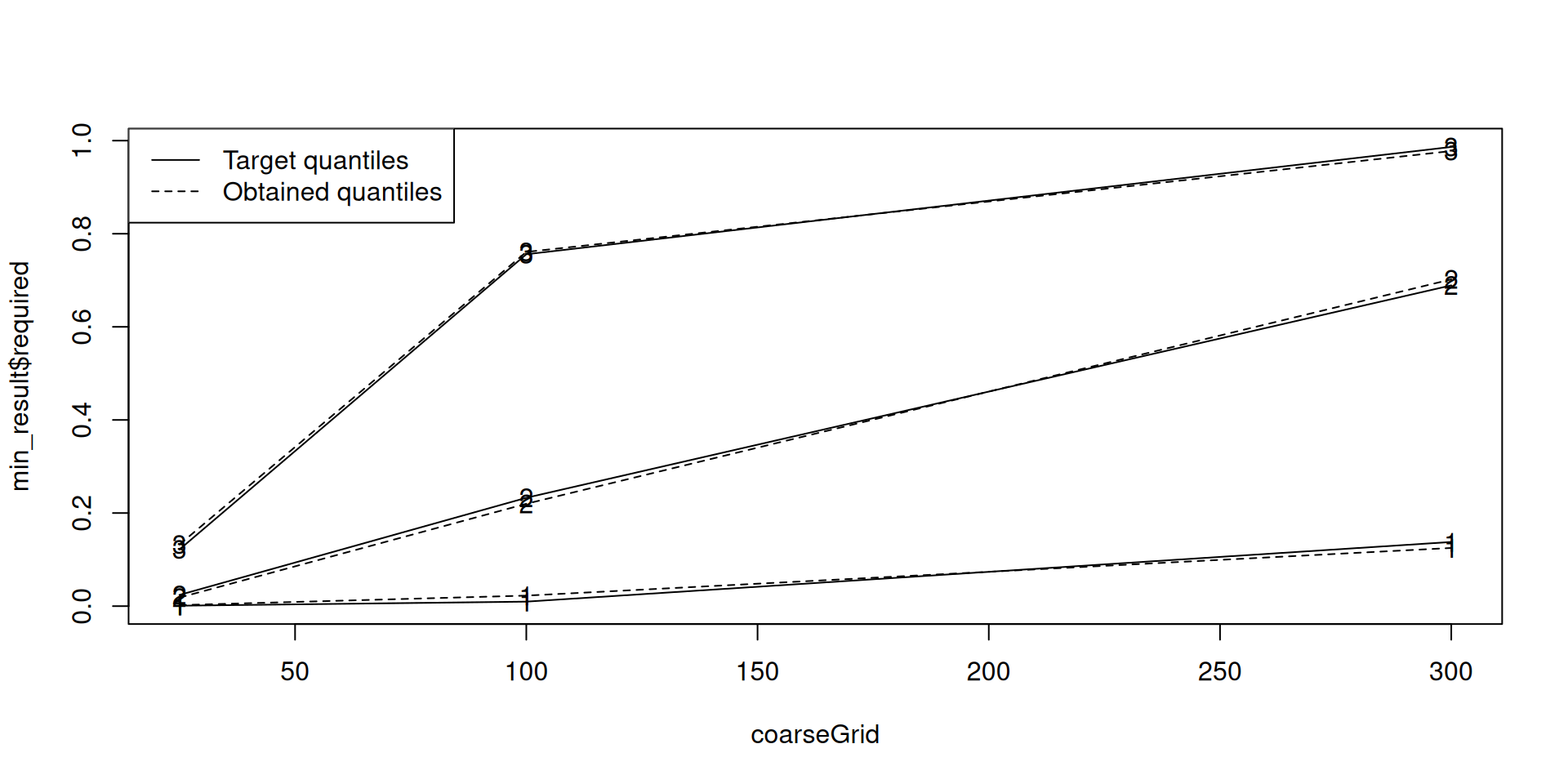
coarseGrid <- c(25, 100, 300)
min_result <- MinimalInformative(
dosegrid = coarseGrid,
refDose = 100,
logNormal = TRUE, # use a log-normal distribution for the slope parameter
threshmin = 0.1, # the quantile for the low dose
threshmax = 0.2, # the quantile for the high dose
seed = 432, # for reproducibility
control = list(max.time = 30) # limit the optimization time here
)It: 1, obj value (lsEnd): 0.06732883077 indTrace: 1
It: 214, obj value (lsEnd): 0.01474025243 indTrace: 214
It: 238, obj value (lsEnd): 0.01416193999 indTrace: 238
It: 460, obj value (lsEnd): 0.01295269405 indTrace: 460
timeSpan = 30.001304 maxTime = 30
Emini is: 0.01295269405
xmini are:
-1.280026342 0.6618977116 1.254935046 0.1500300078 0.5608228116
Totally it used 30.00134 secs
No. of function call is: 10164